5. statsmodels
statsmodel is another statistical library you may use to get more information on your regression models. Let’s take a look at how to do some simple things with this API. Below, we create functions to get data for regression and classification.
[1]:
%matplotlib inline
import matplotlib.pyplot as plt
from sklearn.datasets import make_regression, make_classification
import numpy as np
import pandas as pd
np.random.seed(37)
plt.style.use('ggplot')
def get_regression_data(n_features=5, n_samples=1000):
X, y = make_regression(**{
'n_samples': n_samples,
'n_features': n_features,
'n_informative': n_features,
'n_targets': 1,
'bias': 5.3,
'random_state': 37
})
data = np.hstack([X, y.reshape(-1, 1)])
cols = [f'x{i}' for i in range(n_features)] + ['y']
return pd.DataFrame(data, columns=cols)
def get_classification_data(n_features=5, n_samples=1000):
X, y = make_classification(**{
'n_samples': n_samples,
'n_features': n_features,
'n_informative': n_features,
'n_redundant': 0,
'n_repeated': 0,
'n_classes': 2,
'n_clusters_per_class': 1,
'random_state': 37
})
data = np.hstack([X, y.reshape(-1, 1)])
cols = [f'x{i}' for i in range(n_features)] + ['y']
return pd.DataFrame(data, columns=cols)
5.1. Ordinary least square
An ordinary least square (OLS) model is created using the OLS() function. Below, the patsy API is used to separate the dataframe using R style equation syntax.
[2]:
import statsmodels.api as sm
from patsy import dmatrices
df = get_regression_data()
y, X = dmatrices('y ~ x0 + x1 + x2 + x3 + x4', data=df, return_type='dataframe')
mod = sm.OLS(y, X)
res = mod.fit()
You can also bypass patsy and use the formula API to define the model.
[3]:
import statsmodels.formula.api as smf
mod = smf.ols(formula='y ~ x0 + x1 + x2 + x3 + x4', data=df)
res = mod.fit()
The summary of the data is available through summary().
[4]:
res.summary()
[4]:
| Dep. Variable: | y | R-squared: | 1.000 |
|---|---|---|---|
| Model: | OLS | Adj. R-squared: | 1.000 |
| Method: | Least Squares | F-statistic: | 7.399e+32 |
| Date: | Tue, 01 Dec 2020 | Prob (F-statistic): | 0.00 |
| Time: | 17:09:02 | Log-Likelihood: | 29657. |
| No. Observations: | 1000 | AIC: | -5.930e+04 |
| Df Residuals: | 994 | BIC: | -5.927e+04 |
| Df Model: | 5 | ||
| Covariance Type: | nonrobust |
| coef | std err | t | P>|t| | [0.025 | 0.975] | |
|---|---|---|---|---|---|---|
| Intercept | 5.3000 | 1.01e-15 | 5.23e+15 | 0.000 | 5.300 | 5.300 |
| x0 | 41.8134 | 1.02e-15 | 4.1e+16 | 0.000 | 41.813 | 41.813 |
| x1 | 25.0388 | 1.08e-15 | 2.33e+16 | 0.000 | 25.039 | 25.039 |
| x2 | 3.2108 | 1.02e-15 | 3.14e+15 | 0.000 | 3.211 | 3.211 |
| x3 | 29.9942 | 1.01e-15 | 2.98e+16 | 0.000 | 29.994 | 29.994 |
| x4 | 25.3606 | 1e-15 | 2.53e+16 | 0.000 | 25.361 | 25.361 |
| Omnibus: | 7.219 | Durbin-Watson: | 1.895 |
|---|---|---|---|
| Prob(Omnibus): | 0.027 | Jarque-Bera (JB): | 8.934 |
| Skew: | 0.086 | Prob(JB): | 0.0115 |
| Kurtosis: | 3.430 | Cond. No. | 1.16 |
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
The params properties of the results will retrieve the coefficients of the model.
[5]:
res.params
[5]:
Intercept 5.300000
x0 41.813417
x1 25.038808
x2 3.210812
x3 29.994205
x4 25.360629
dtype: float64
The rsquared property will retrieve the \(R^2\) value.
[6]:
res.rsquared
[6]:
1.0
There are a lot of model diagnostic functions available. Below, we apply the rainbow test for linearity.
[7]:
f_stats, p_value = sm.stats.linear_rainbow(res)
print(f'f-statistic: {f_stats:.5f}, p-value: {p_value:.5f}')
f-statistic: 0.24804, p-value: 1.00000
5.2. Logistic regression
Logistic regression is a type of generalized linear model. A logistic regression model is created using the GLM() function. Note that y comes before X for GLM(), as opposed to X coming before y when using OLS().
[8]:
df = get_classification_data()
y, X = dmatrices('y ~ x0 + x1 + x2 + x3 + x4', data=df, return_type='dataframe')
binonmial_model = sm.GLM(y, X, family=sm.families.Binomial())
res = binonmial_model.fit()
[9]:
res.summary()
[9]:
| Dep. Variable: | y | No. Observations: | 1000 |
|---|---|---|---|
| Model: | GLM | Df Residuals: | 994 |
| Model Family: | Binomial | Df Model: | 5 |
| Link Function: | logit | Scale: | 1.0000 |
| Method: | IRLS | Log-Likelihood: | -76.314 |
| Date: | Tue, 01 Dec 2020 | Deviance: | 152.63 |
| Time: | 17:09:02 | Pearson chi2: | 9.15e+05 |
| No. Iterations: | 9 | ||
| Covariance Type: | nonrobust |
| coef | std err | z | P>|z| | [0.025 | 0.975] | |
|---|---|---|---|---|---|---|
| Intercept | -1.5205 | 0.636 | -2.389 | 0.017 | -2.768 | -0.273 |
| x0 | 2.2090 | 0.259 | 8.522 | 0.000 | 1.701 | 2.717 |
| x1 | -3.1632 | 0.348 | -9.083 | 0.000 | -3.846 | -2.481 |
| x2 | 2.0077 | 0.248 | 8.095 | 0.000 | 1.522 | 2.494 |
| x3 | -1.0259 | 0.301 | -3.408 | 0.001 | -1.616 | -0.436 |
| x4 | 2.6748 | 0.382 | 7.010 | 0.000 | 1.927 | 3.423 |
5.3. ANOVA
Analysis of Variance (ANOVA) may be conducted. The data below is taken from here.
[10]:
df = pd.DataFrame({
'A': [25, 30, 28, 36, 29],
'B': [45, 55, 29, 56, 40],
'C': [30, 29, 33, 37, 27],
'D': [54, 60, 51, 62, 73]
})
df
[10]:
| A | B | C | D | |
|---|---|---|---|---|
| 0 | 25 | 45 | 30 | 54 |
| 1 | 30 | 55 | 29 | 60 |
| 2 | 28 | 29 | 33 | 51 |
| 3 | 36 | 56 | 37 | 62 |
| 4 | 29 | 40 | 27 | 73 |
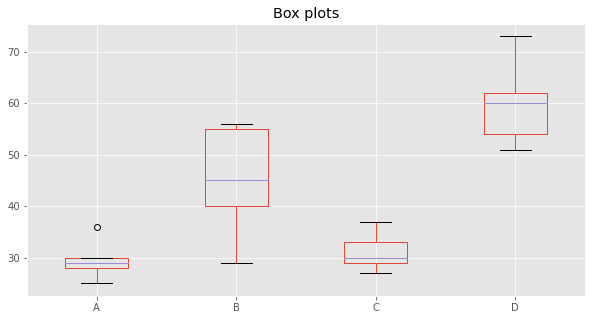
Let’s do some box plots.
[11]:
fig, ax = plt.subplots(figsize=(10, 5))
_ = df.plot(kind='box', ax=ax)
_ = ax.set_title('Box plots')
scipy can do the one-way ANOVA.
[12]:
import scipy.stats as stats
f, p = stats.f_oneway(*[df[c] for c in df.columns])
print(f'f-statistics: {f:.5f}, p-value: {p:.5f}')
f-statistics: 17.49281, p-value: 0.00003
Before statsmodel can do ANOVA, we have to melt the dataframe.
[13]:
ldf = pd.melt(df.reset_index(), id_vars=['index'], value_vars=['A', 'B', 'C', 'D'])\
.rename(columns={'index': 'index', 'variable': 'treatment', 'value': 'value'})
ldf.head()
[13]:
| index | treatment | value | |
|---|---|---|---|
| 0 | 0 | A | 25 |
| 1 | 1 | A | 30 |
| 2 | 2 | A | 28 |
| 3 | 3 | A | 36 |
| 4 | 4 | A | 29 |
Now we specify the ANOVA model and acquire the table.
[14]:
anova_model = smf.ols('value ~ C(treatment)', data=ldf).fit()
anova_table = sm.stats.anova_lm(anova_model, typ=2)
anova_table
[14]:
| sum_sq | df | F | PR(>F) | |
|---|---|---|---|---|
| C(treatment) | 3010.95 | 3.0 | 17.49281 | 0.000026 |
| Residual | 918.00 | 16.0 | NaN | NaN |
Post-hoc testing is also possible.
[15]:
from statsmodels.stats.multicomp import pairwise_tukeyhsd
res = pairwise_tukeyhsd(ldf.value, ldf.treatment)
res.summary()
[15]:
| group1 | group2 | meandiff | p-adj | lower | upper | reject |
|---|---|---|---|---|---|---|
| A | B | 15.4 | 0.0251 | 1.6929 | 29.1071 | True |
| A | C | 1.6 | 0.9 | -12.1071 | 15.3071 | False |
| A | D | 30.4 | 0.001 | 16.6929 | 44.1071 | True |
| B | C | -13.8 | 0.0482 | -27.5071 | -0.0929 | True |
| B | D | 15.0 | 0.0296 | 1.2929 | 28.7071 | True |
| C | D | 28.8 | 0.001 | 15.0929 | 42.5071 | True |